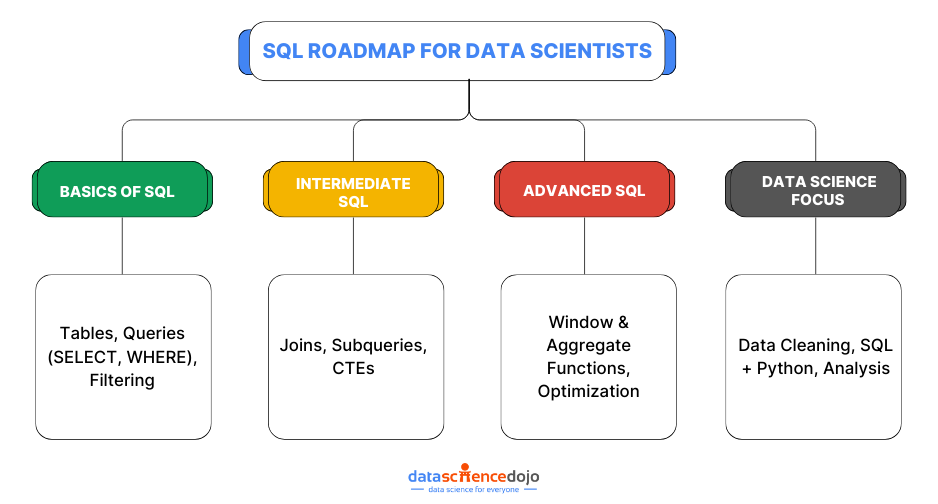
SQL for data scientists is more than just a querying tool-it’s a critical skill for extracting, transforming, and analyzing structured data efficiently. Mastering SQL allows data scientists to efficiently process large datasets, uncover patterns, and make informed decisions based on their findings.
At the core of SQL proficiency is a strong understanding of its syntax. Essential commands such as SELECT, WHERE, JOIN, and GROUP BY enable users to filter, aggregate, and organize data with precision. These statements form the backbone of SQL operations, allowing data scientists to perform everything from simple lookups to complex data transformations.
Equally important is understanding how data is structured within relational databases. Relationships such as one-to-one, one-to-many, and many-to-many dictate how tables interact, and knowing how to work with foreign keys, joins, and normalization techniques ensures data integrity and efficient retrieval. Without this knowledge, querying large datasets can become inefficient and error-prone.
This blog delves into 12 essential SQL concepts that every data scientist should master. Through real-world examples and best practices, it will help you write efficient, scalable queries—whether you’re just starting out or looking to refine your SQL expertise.
Here’s an interesting read about Top 10 SQL commands
Let’s dive into some of the key SQL concepts that are important to learn for a data scientist.
1. Formatting Strings
Cleaning raw data is essential for accurate analysis and improved decision-making. String functions provide powerful tools to manipulate and standardize text, ensuring consistency across datasets.
The CONCAT function merges multiple strings into a single value, making it useful for formatting names, addresses, or reports. Handling missing values efficiently, COALESCE replaces NULL entries with predefined defaults, preventing data gaps and ensuring completeness. Leveraging these functions enhances readability, maintains data integrity, and boosts overall productivity.
2. Stored Methods
Stored procedures are precompiled collections of SQL statements that can be executed as a single unit, improving performance, reusability, and maintainability.
They optimize performance by reducing execution time, as they are stored and compiled in the database, minimizing network traffic. Reusability ensures that complex queries don’t need to be rewritten, and any updates to the procedure apply universally. Security is enhanced by allowing controlled access to data while reducing injection risks. Stored procedures also encapsulate business logic, making database operations more structured and manageable.
Modifications can be made using ALTER PROCEDURE, and procedures can be removed with DROP PROCEDURE. Overall, stored procedures streamline database operations by reducing redundancy, improving efficiency, and centralizing logic, making them essential for scalable database management.
3. Joins
Joins in SQL allow you to combine data from multiple tables based on defined relationships, making data retrieval more efficient and meaningful. An INNER JOIN returns only the matching records from both tables, functioning like the intersection of two sets. This ensures that only relevant data common to both tables is retrieved.
A LEFT JOIN returns all records from the left table and only matching records from the right table. If no match exists, the result still includes records from the left table with NULL values for missing data from the right table. Conversely, a RIGHT JOIN includes all records from the right table and only matching records from the left table, filling unmatched left-side records with NULL values.
Understanding these joins is crucial for accurate data extraction, preventing unnecessary clutter while ensuring that the right relationships between tables are utilized.
4. Subqueries
A subquery is a query within another query, allowing for structured data filtering and processing. It is especially useful when working with multiple tables or when intermediate computations are needed before executing the main query. Subqueries help break down complex queries into manageable steps, improving readability and efficiency.
When a subquery returns a single value, it can be used directly in conditions like comparisons. However, if a subquery returns multiple rows, multi-line operators like IN or EXISTS are required to handle the results properly. These operators ensure that the main query processes multiple values correctly without errors. Understanding subqueries enhances query flexibility, enabling more dynamic and precise data retrieval.
5. Normalization
Normalization is a fundamental SQL concept because it directly impacts database design and query performance. SQL databases use normalization techniques to structure tables efficiently, reducing redundancy and improving data integrity. When designing a relational database, SQL statements like CREATE TABLE, FOREIGN KEY, and JOIN work based on the principles of normalization.
For example, when you normalize a database, you often break large, redundant tables into smaller ones and use foreign keys to maintain relationships. This affects how SQL queries are written, especially in SELECT, INSERT, and UPDATE operations.
Well-normalized databases lead to optimized JOIN performance and prevent anomalies that could corrupt data integrity. Thus, normalization is not just a theoretical concept but a practical SQL design strategy essential for creating efficient and scalable databases.
Another interesting read: SQL vs NoSQL
6. Manipulating Dates and Times
Manipulating Dates and Times in SQL is essential for organizing and analyzing time-based data efficiently. SQL provides various functions to extract, calculate, and modify date values based on specific requirements.
The EXTRACT function allows you to pull specific components such as year, month, or day from a date, making it easier to categorize and filter data. The DATEDIFF function calculates the difference between two dates, which is useful for measuring durations like age, time between events, or project deadlines.
Additionally, DATE_ADD and DATE_SUB allow you to shift dates forward or backward by a specified number of days, months, or years, making it easy to adjust time-based data dynamically.
These date functions help in organizing data chronologically, facilitating trend analysis, and ensuring accurate time-based reporting.
7. Transactions
A transaction in SQL is a sequence of operations executed as a single unit of work to ensure data integrity and consistency. Transactions follow the ACID properties: Atomicity (all operations complete or none at all), Consistency (data remains valid before and after the transaction), Isolation (concurrent transactions do not interfere with each other), and Durability (changes are permanently saved once committed).
Key commands include BEGIN TRANSACTION to start a transaction, COMMIT to save changes, and ROLLBACK to undo changes if an error occurs. Transactions are essential in scenarios like banking, where money must be deducted from one account and added to another—if one step fails, the entire transaction is rolled back to prevent data inconsistencies.
8. Connecting SQL to Python or R
SQL is powerful for managing and querying databases, but integrating it with Python or R unlocks advanced data analysis, machine learning, and visualization capabilities. By using libraries like pandas and sqlite3 in Python or dplyr and DBI in R, you can seamlessly extract, manipulate, and analyze SQL data within a coding environment.
Python’s pandas allows direct SQL queries with functions like read_sql(), making it easy to transform data for machine learning models. Similarly, R’s dplyr simplifies SQL queries while offering extensive statistical and visualization tools. Mastering SQL integration with these languages enhances workflow efficiency and is essential for data science, automation, and business intelligence applications.
You might also like: SnowSQL
9. Features of Window Functions
Window functions enable calculations across a set of rows while preserving individual row details. Unlike aggregate functions that collapse data into a single result, window functions retain row-level granularity while applying computations over a defined window.
The OVER clause determines how the window is structured, using PARTITION BY to group data into subsets and ORDER BY to establish sorting within each partition. Common applications include RANK for ranking rows, LAG and LEAD for accessing previous or next values, and moving averages for trend analysis. These functions are essential for advanced analytical queries, providing deeper insights without losing row-specific details.
10. Indexing for Performance Optimization
Indexes enhance query performance by enabling faster data retrieval. Instead of scanning entire tables, an index helps locate specific rows more efficiently, reducing execution time for searches and lookups.
Applying indexes to frequently queried columns can significantly speed up operations, especially in large datasets. However, excessive indexing can negatively impact performance by slowing down insertions, updates, and deletions, as each modification requires updating the associated indexes. Striking a balance between fast retrieval and efficient data manipulation is essential for optimal performance.
11. Predicates
Predicates, used in WHERE, HAVING, and JOIN clauses, refine data selection by filtering records before processing. Applying precise predicates minimizes the number of rows scanned, improving query performance and reducing computational costs.
Using conditions like filtering by specific dates, ranges, or categories ensures only relevant data is retrieved. For example, restricting results to today’s signups with a date filter significantly reduces processing time, which is especially beneficial in cloud-based environments where query efficiency directly impacts costs. Effective use of predicates enhances both speed and resource management.
12. Query Syntax
Structured query syntax enables efficient data retrieval by following a logical sequence. Every query begins with SELECT to choose columns, FROM to specify tables, and WHERE to apply filters, ensuring only relevant data is processed.
Understanding how these clauses interact allows for writing optimized queries that balance performance and readability. Mastering structured query syntax streamlines data extraction, making analysis more intuitive while improving efficiency in handling large datasets.
Here’s a list of Techniques for Data Scientists to Upskill with LLMs
SQL for Data Scientists – A Must-Have Skill
Mastering SQL for data scientists is essential for efficiently querying, managing, and analyzing structured data. From understanding basic syntax to optimizing complex queries and handling database relationships, SQL plays a crucial role in extracting meaningful insights. By honing these skills, data scientists can work more effectively with large datasets, improve decision-making, and enhance their overall analytical capabilities.
Whether you’re just starting out or looking to refine your expertise, a strong foundation in SQL will always be a valuable asset in the world of data science.